

injurytools is a package designed for the field of Sports Medicine. It simplifies the data analysis workflow by providing convenience functions and handy tools for sports injury data.
The functions can be classified into: (a) sports injury data preparation, (b) descriptive analyses and (c) data visualization routines. Further analyses, such as the estimation of the risk of injury with other covariate effects, can be performed outside of injurytools, whether the event of injury is viewed as count or time-to-event data.
To get an overview of the package, see the Articles section.
In practice, the package can help automate certain descriptive reports that are routinely made for sports injury surveillance.
To install from CRAN:
To install the most current version from GitHub:
Most functions contain or start with inj*() which stands for injury. Functions for data preparation start with prepare_*(); and those for data visualization with gg_inj*().
The below outlines at a glance how injurytools can help to get a comprehensive picture of injury data:
library(injurytools)
library(ggplot2)
gg_injphoto(injd,
title = "Overview of injuries:\nLiverpool FC 1st male team during 2017-2018 and 2018-2019 seasons",
by_date = "2 month") +
## plus some lines of ggplot2 code..
xlab("Follow-up date") + ylab("Players") +
labs(caption = "source: transfermarkt.com") +
theme(plot.title = element_text(face = "bold", hjust = 0.5, size = 22),
axis.text.x.bottom = element_text(size = 13, angle = 20, hjust = 1),
axis.text.y.left = element_text(size = 12),
axis.title.x = element_text(size = 20, face = "bold", vjust = -1),
axis.title.y = element_text(size = 20, face = "bold", vjust = 1.8),
legend.text = element_text(size = 20),
plot.caption = element_text(face = "italic", size = 12, colour = "gray10"))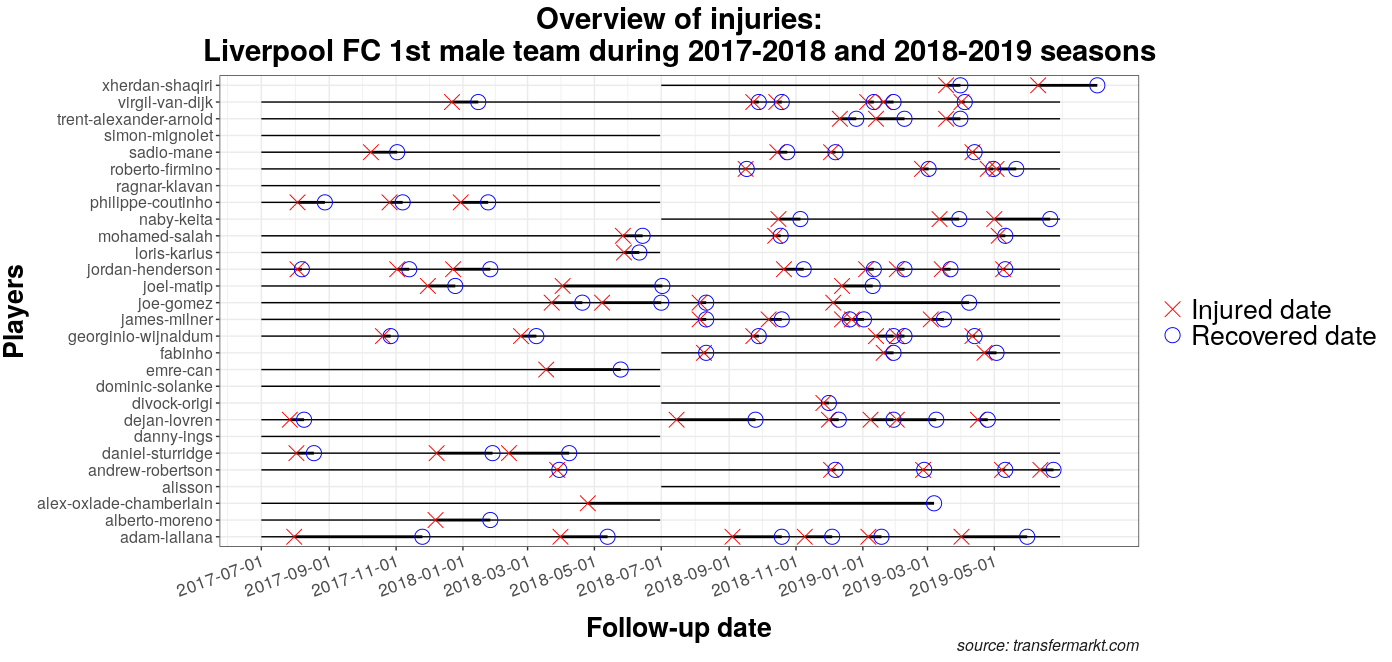
If you have problems with the package, find any bugs, or have suggestions for improvements, please feel free to open a GitHub issue or touch us directly via email. We also welcome your feedback.