

The main content of pedprobr is an implementation of the Elston-Stewart algorithm for pedigree likelihoods given marker genotypes. It is part of the pedsuite, a collection of packages for pedigree analysis in R.
The pedprobr package does much of the hard work in several other pedsuite packages:
The workhorse of pedprobr is the likelihood() function, supporting a variety of situations:
To get the current official version of pedprobr, install from CRAN as follows:
Alternatively, you can obtain the latest development version from GitHub:
# install.packages("devtools") # install devtools if needed
devtools::install_github("magnusdv/pedprobr")To set up a simple example, we first use pedtools utilities to create a pedigree where two brothers are genotyped with a single SNP marker. The marker has alleles a and b, with frequencies 0.2 and 0.8 respectively, and both brothers are heterozygous a/b.
# Pedigree with SNP marker
x = nuclearPed(nch = 2) |>
addMarker(geno = c(NA, NA, "a/b", "a/b"), afreq = c(a = 0.2, b = 0.8))
# Plot with genotypes
plot(x, marker = 1)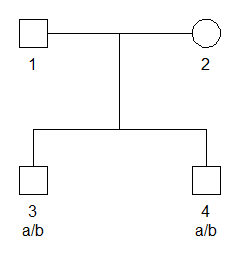
The pedigree likelihood, i.e., the probability of the genotypes given the pedigree, is obtained as follows:
Besides likelihood(), other important functions in pedprobr are:
oneMarkerDistribution() : the joint genotype distribution at a single marker, for any subset of pedigree memberstwoMarkerDistribution() : the joint genotype distribution at two linked markers, for a single personIn both cases, the distributions are computed conditionally on any known genotypes at the markers in question.
To illustrate oneMarkerDistribution() we continue our example from above, and consider the following question: What is the joint genotype distribution of the parents, conditional on the genotypes of the children?
The answer is found as follows:
oneMarkerDistribution(x, ids = 1:2, partialmarker = 1, verbose = F)
#> a/a a/b b/b
#> a/a 0.00000000 0.01724138 0.1379310
#> a/b 0.01724138 0.13793103 0.2758621
#> b/b 0.13793103 0.27586207 0.0000000The output confirms the intuitive result that the parents cannot both be homozygous for the same allele. The most likely combination is that one parent is heterozygous a/b, while the other is homozygous b/b.