

{shapviz} provides typical SHAP plots:
sv_importance(): Importance plots (bar plots and/or beeswarm plots).sv_dependence() and sv_dependence2D(): Dependence plots to study feature effects and interactions.sv_interaction(): Interaction plots.sv_waterfall(): Waterfall plots to study single predictions.sv_force(): Force plots as alternative to waterfall plots.SHAP and feature values are stored in a “shapviz” object that is built from:
# From CRAN
install.packages("shapviz")
# Or the newest version from GitHub:
# install.packages("devtools")
devtools::install_github("ModelOriented/shapviz")Shiny diamonds… let’s use XGBoost to model their prices by the four “C” variables:
library(shapviz)
library(ggplot2)
library(xgboost)
set.seed(1)
# Build model
x <- c("carat", "cut", "color", "clarity")
dtrain <- xgb.DMatrix(data.matrix(diamonds[x]), label = diamonds$price)
fit <- xgb.train(params = list(learning_rate = 0.1), data = dtrain, nrounds = 65)
# SHAP analysis: X can even contain factors
dia_2000 <- diamonds[sample(nrow(diamonds), 2000), x]
shp <- shapviz(fit, X_pred = data.matrix(dia_2000), X = dia_2000)
sv_importance(shp, show_numbers = TRUE)
sv_dependence(shp, v = x)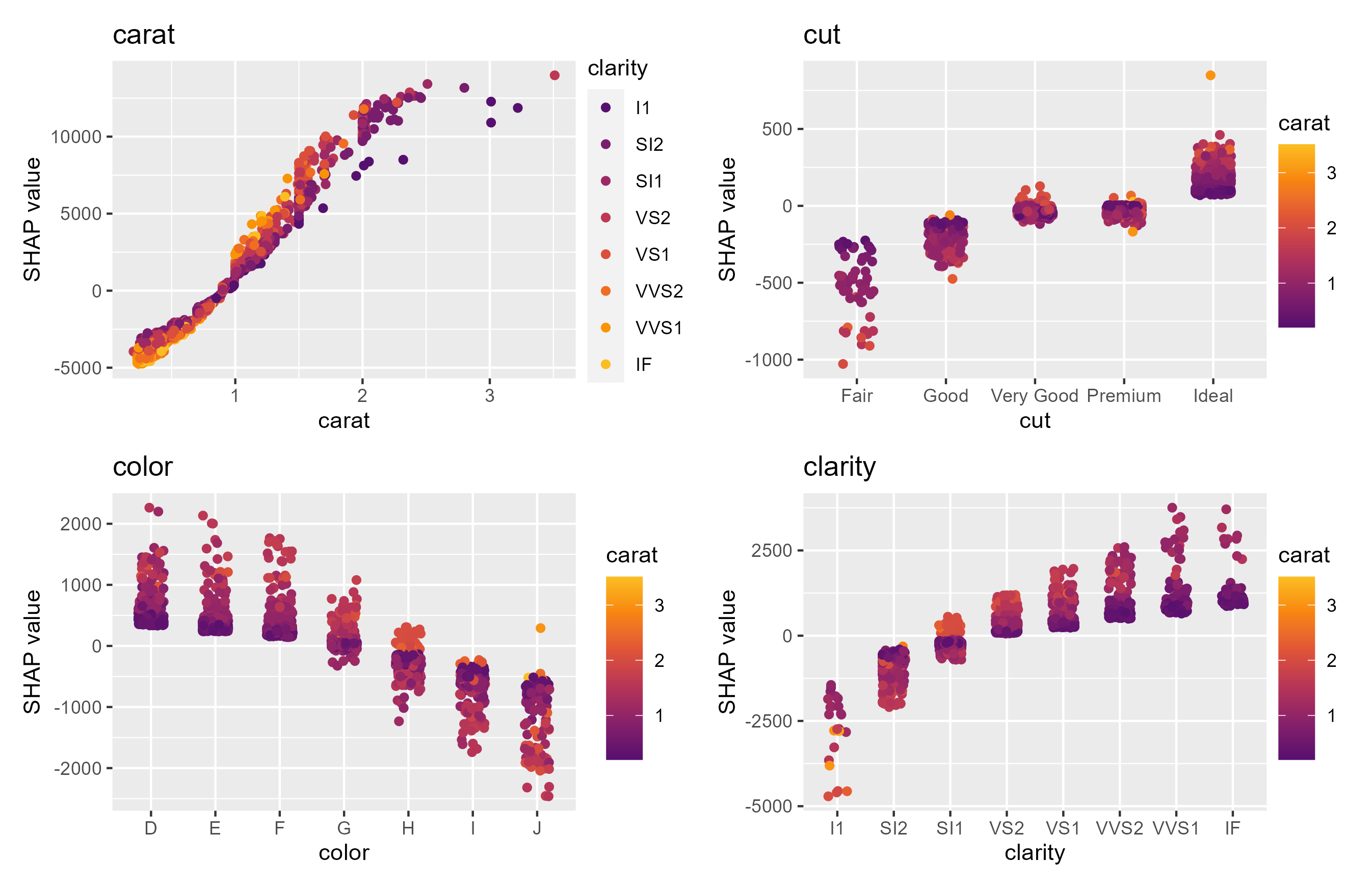
Decompositions of individual predictions can be visualized as waterfall or force plot:
Check-out the vignettes for topics like:
[1] Scott M. Lundberg and Su-In Lee. A Unified Approach to Interpreting Model Predictions. Advances in Neural Information Processing Systems 30 (2017).